Research Overview
Even genetically identical bacterial cells grown under the same conditions often express different sets of genes and behave differently. Not unlike individual cells in the human body, they may assume specialized roles for the benefit of the whole community. We are developing and applying high-throughput single-cell genomic technologies to identify and characterize diverse functional subpopulations of bacteria within complex microbial samples. We are interested in unraveling the roles and behavior of individual bacteria in challenging environments such as the human host, and within structured multi-species communities known as biofilms.

Biofilm gene expression
Biofilms are spatially structured bacterial communities that contribute to chronic bacterial diseases. Many cells within the biofilm are shielded from host defenses and drugs.
Host-pathogen interactions
Even in a lab-grown monoculture, bacterial gene expression can be highly heterogeneous. Many mammalian cell types also exhibit considerable variability in gene expression states.
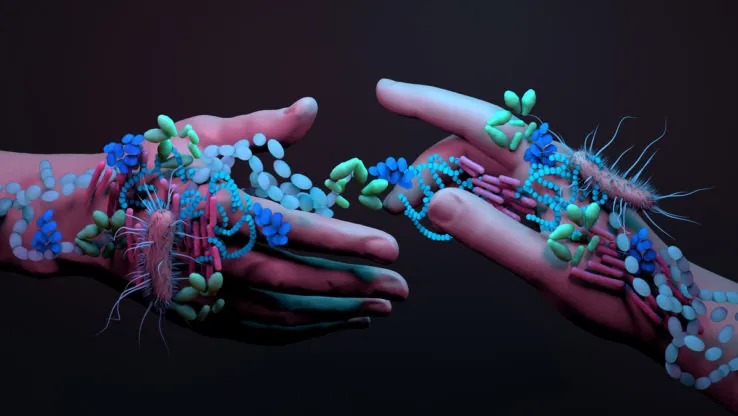

Single-cell biology of microbiome
The gut microbiota is currently understood to represent a complex “organ” in the body affecting a multitude of body systems and implicated in a wide range of diseases.
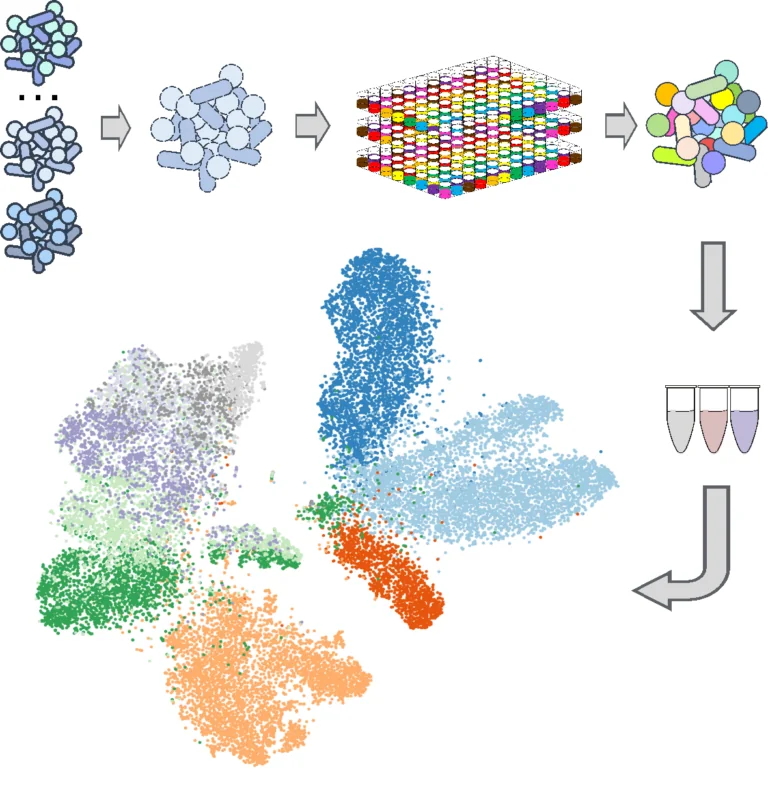
Biotechnology Development
We are passionate about developing, improving and repurposing high-throughput technologies for gaining insight into bacterial communities.